
Depiction of the samurai Suenaga Takezaki repelling Mongol and Korean arrows and bombs at Torikai-Gata. “Mōko Shūrai Ekotoba” by 竹崎季長 – 蒙古襲来絵詞. Open source: Wikimedia Commons
Genetics researchers in a 2003 report, The Genetic Legacy of the Mongols, AJHG, Volume 72, Issue 3, March 2003, Pages 717–721, Tatiana Zerjal, et al., found that the Mongol Y-DNA marker showed up in 16 populations throughout a large region of Asia, stretching from the Pacific to the Caspian Sea. The report also identified the origin to be “most likely in Mongolia, where the largest number of different star-cluster haplotypes is found ( fig. 1). Thus, a single male line, probably originating in Mongolia, has spread in the last ∼1,000 years to represent ∼8% of the males in a region stretching from northeast China to Uzbekistan”.
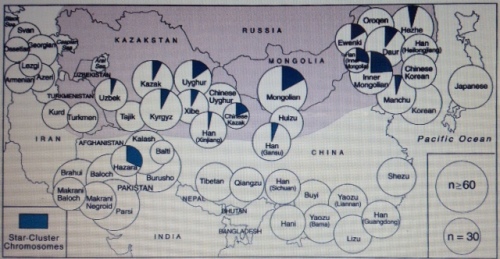
Researchers show the footprint of Genghis Khan Y-chromosome star-cluster marker across Eurasia. The Mongols left no genetic trace in Japan, however, corroborating the historical accounts of the Japanese having successfully repelled the Mongols and eventually driving them back to the mainland, despite the earlier devastating defeats during the Battles of Bunei. Fig. 2 from the report
Excerpted from the report:
“We have identified a Y-chromosomal lineage with several unusual features. It was found in 16 populations throughout a large region of Asia, stretching from the Pacific to the Caspian Sea, and was present at high frequency: ∼8% of the men in this region carry it, and it thus makes up ∼0.5% of the world total. The pattern of variation within the lineage suggested that it originated in Mongolia ∼1,000 years ago. Such a rapid spread cannot have occurred by chance; it must have been a result of selection. The lineage is carried by likely male-line descendants of Genghis Khan, and we therefore propose that it has spread by a novel form of social selection resulting from their behavior.
The patterns of variation found in human DNA are usually considered to result from a balance between neutral processes and natural selection. Among the former, mutation, recombination, and migration increase variation, whereas genetic drift decreases it. Natural selection can act to remove deleterious variants (purifying selection), maintain polymorphism (balancing selection), or produce a trend (directional selection). Clear examples of the latter are rare in humans, but probable cases, such as those associated with resistance to malaria (Hamblin and Di Rienzo 2000) or unidentified pathogens (Stephens et al. 1998), can be recognized by the “signature” they leave in the genome. The rapid increase in frequency of the selected allele and its linked sequences results in a haplotype that is found at higher frequency than would be expected from its degree of variation. We have now identified such a haplotype on the Y chromosome, but we suggest that its spread results not from a biological advantage, but from human activities recorded in history.
In surveys of DNA variation in Asia, we typed 2,123 men with 32 markers to produce a Y haplotype for each man; these included 1,126 individuals described elsewhere (Qamar et al. 2002; Zerjal et al. 2002). Over 90% of the haplotypes showed the usual pattern (Mohyuddin et al. 2001): most males had a unique code; and the few haplotypes present in more than one individual were generally found within the same population. However, we also saw one pattern that was novel in two respects. First, there was a high frequency of a cluster of closely related lineages, collectively called the “star cluster” (fig. 1, shaded area). Second, star-cluster chromosomes were found in 16 populations throughout a large geographical area extending from Central Asia to the Pacific ( fig. 2); thus, they do not result from an event specific to any single population. We can deduce the most likely time to the most recent common ancestor (TMRCA) and place of origin of this unusual lineage from the observed genetic variation. To do this, it is first necessary to distinguish star-cluster chromosomes from the remainder. For this, we used the criterion that haplotypes linked to the central one in the shaded area of the network without gaps would be included ( fig. 1). We then used two approaches to calculate a TMRCA for the star-cluster chromosomes. The program BATWING (Wilson and Balding 1998) uses models of both mutation and population processes, which were specified as described elsewhere (Qamar et al. 2002). With this program, we estimated ∼1,000 years for the TMRCA (95% confidence interval limits ∼700–1,300 years). The use of alternative demographic models with constant or exponentially increasing population size changed the estimate by <10%. A method that does not consider population structure (Morral et al. 1994), ρ, suggested ∼860 (∼590–1,300) years. In both calculations, we assumed a generation time of 30 years. The origin was most likely in Mongolia, where the largest number of different star-cluster haplotypes is found ( fig. 1). Thus, a single male line, probably originating in Mongolia, has spread in the last ∼1,000 years to represent ∼8% of the males in a region stretching from northeast China to Uzbekistan. If this spread were due to a general population expansion, we would expect to find multiple lineages with the same characteristics of high frequency and presence in multiple populations, but we do not (Zerjal et al. 2002). The star-cluster pattern is unique.”
Was it due to selection?
“This rise in frequency, if spread evenly over ∼34 generations, would require an average increase by a factor of ∼1.36 per generation and is thus comparable to the most extreme selective events observed in natural populations, such as the spread of melanic moths in 19thcentury England in response to industrial pollution (Edleston 1865). We evaluated whether it could have occurred by chance. If the population growth rate is known, it is possible to test whether the observed frequency of a lineage is consistent with its level of variation, assuming neutrality (Slatkin and Bertorelle 2001). Using this method, we estimated the chance of finding the low degree of variation observed in the star cluster, with a current frequency of ∼8%, under neutral conditions. Even with the demographic model most likely to lead to rapid increase of the lineage, double exponential growth, the probability was !10237; if the mutation rate were 10 times lower, the probability would still be !1010. Thus, chance can be excluded: selection must have acted on this haplotype. Could biological selection be responsible? Although this possibility cannot be entirely ruled out, the small number of genes on the Y chromosome and their specialized functions provide few opportunities for selection (Jobling and Tyler-Smith 2000). It is therefore necessary to look for alternative explanations. Increased reproductive fitness, transmitted socially from generation to generation, of males carrying the same Y chromosome would lead to the increase in frequency of their Y lineage, and this effect would be enhanced by the elimination of unrelated males. Within the last 1,000 years in this part of the world, these conditions are met by Genghis (Chingis) Khan (c. 1162–1227) and his male relatives. He established the largest land empire in history and often slaughtered the conquered populations, and he and his close male relatives had many children. Although the Mongol empire soon disintegrated as a political unit, his male-line descendants ruled large areas of Asia for many generations. These included China, where the Yuan Dynasty emperors remained in power until 1368, after which the Mongols continued to dominate the country north of the Great Wall for several more centuries, and the region west to the Aral Sea, where the Chaghatai Khans ruled. Although their power diminished over time, they remained at Kashghar near the Kyrgyzstan/ China border until the middle of the 17th century (Morgan 1986). It is striking that the boundary of the Mongol empire when Genghis Khan died (fig. 2), which also corresponds to the boundaries of the regions controlled by later Khans, matches the distribution of star-cluster chromosomes closely, with one exception: the Hazaras. We, therefore, wished to compare Genghis Khan’s Y profile with the star cluster. It is not possible to examine his remains directly, but history provides an alternative. The Hazaras of Pakistan have a Mongol origin (Qamar et al. 2002), and many consider themselves to be direct male-line descendants of Genghis Khan. A genealogy documenting these links has been constructed from their oral history (Mousavi 1998). A large proportion of the Hazara pro- files do indeed lie in the star cluster, which is not otherwise seen in Pakistan (fig. 2), thus supporting their oral tradition and suggesting that Genghis Khan carried the star-cluster haplotype. The Y chromosome of a single individual has spread rapidly and is now found in ∼8% of the males throughout a large part of Asia. Indeed, if our sample is representative, this chromosome will be present in about 16 million men, ∼0.5% of the world’s total. The available evidence suggests that it was carried by Genghis Khan. His Y chromosome would obviously have had ancestors, and our best estimate of the TMRCA of starcluster chromosomes lies several generations before his birth. … The historically documented events accompanying the establishment of the Mongol empire would have contributed directly to the spread of this lineage by Genghis Khan and his relatives, but perhaps as important was the establishment of a long-lasting male dynasty
See the rest of the report here.
Read also:
Around 500 confirmed sunken shipwreck sites in Japanese waters lack funding for underwater archaeological survey and research work (Heritage of Japan, this site)
Sex and power. The reproductive instinct of conquerors (Social Ethology)
The Mongol Invasions of Korea | The Mongol Invasions of Japan
The Mongol Invasions 1274 and 1281, by Stephen Turnbull, Osprey Publishing 2010
